Contest tasks

Task 1.
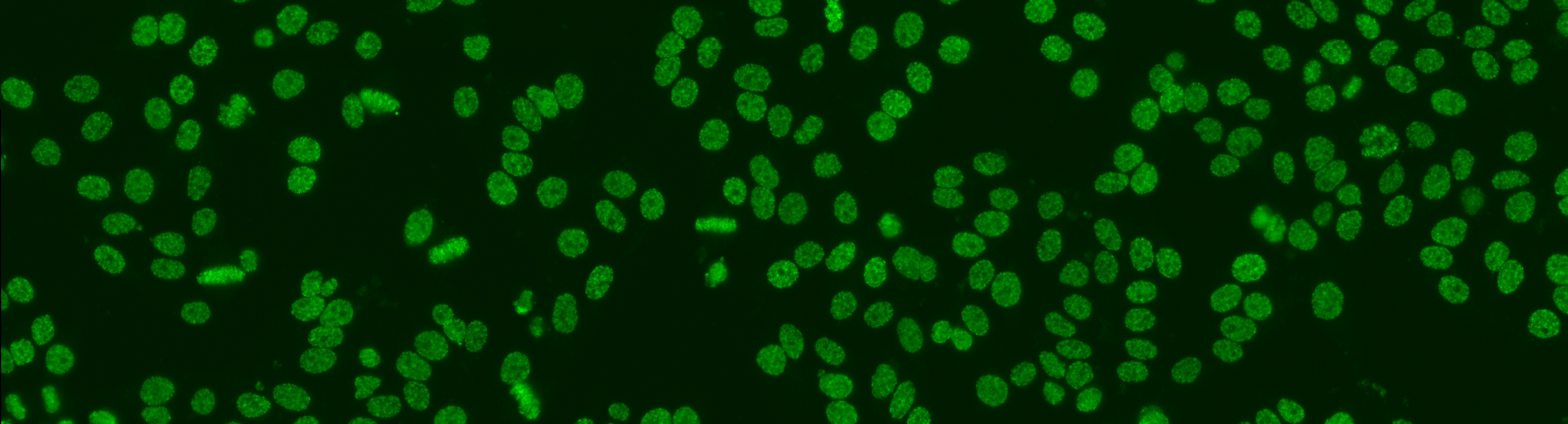
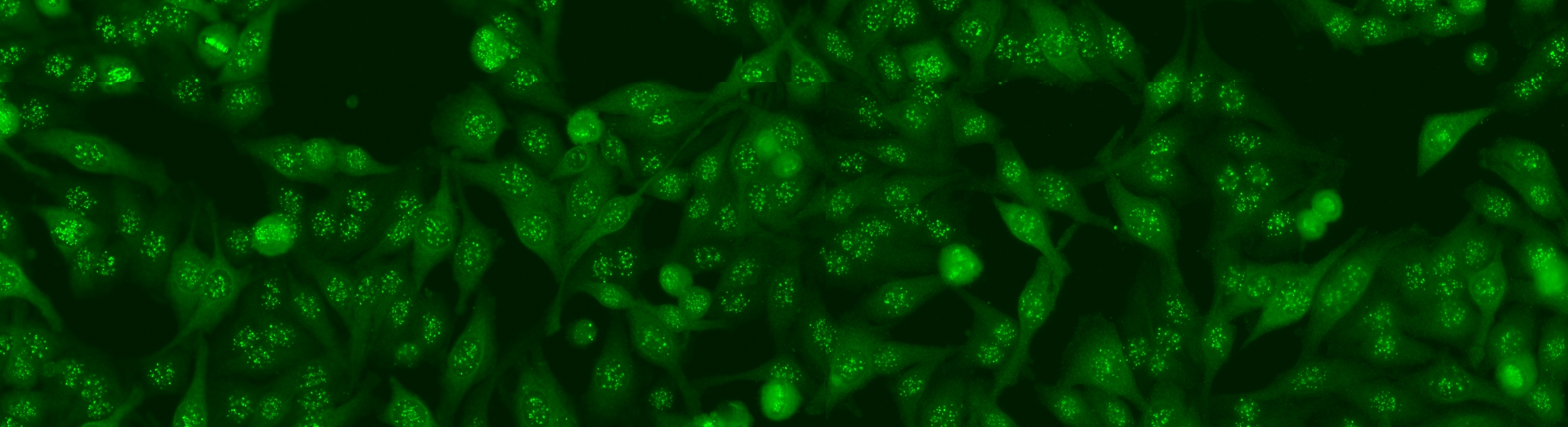
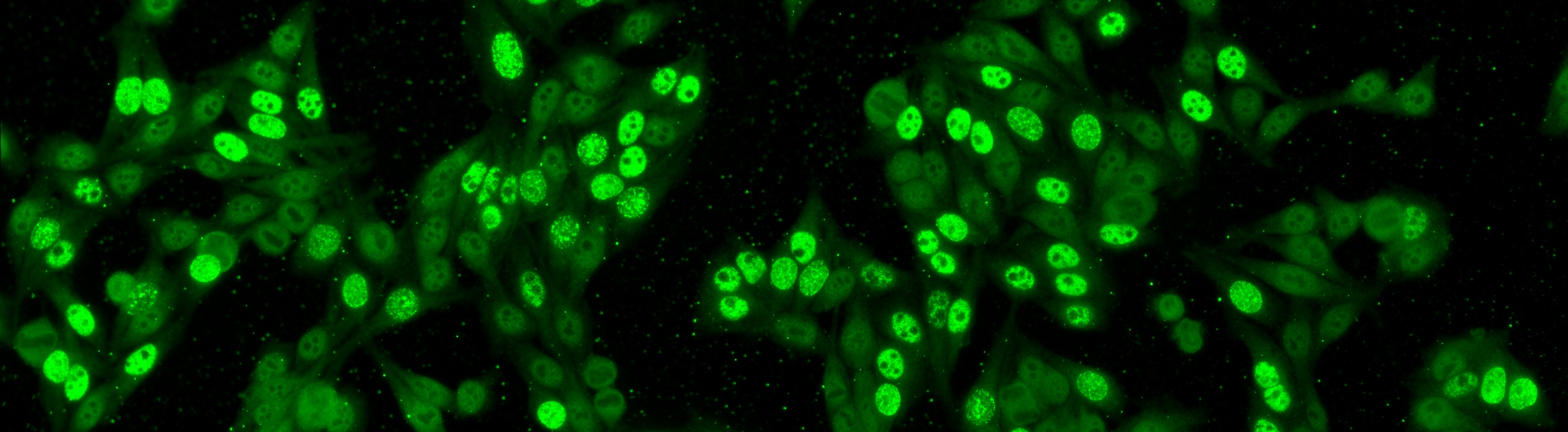
Similar to the contest from the previous year, this task is focused on cell-level pattern classification. This task has been initially proposed at the “HEp-2 Cells Classification” contest hosted by the ICPR 2012 and then proposed again in all the successive initiatives hosted by the ICIP 2013 and ICPR 2014, using datasets of increased size and complexities of the patterns. In particular, the competitions hold on years 2013 and 2014 considered the following six HEp-2 patterns: homogeneous, centromere, speckled, nucleolar, mitotic spindle and Golgi. This task will be the continuation from the previous year to witness the advances made by the community. More info.
Task 2.
The aim of the task is to classify the patient specimen. Each patient specimen (refer to Fig. 4) have multiple HEp-2 cells. This task will equip the participants with the segmented HEp-2 cells although the mitotic cells are not indicated. The participants have the liberty to either work directly on the specimen images (i.e. without using the segmentation map), or utilize the provided segmentation map. This task has been firstly proposed at the ICPR 2014 competition and similarly to the cell-level pattern classification task. Again, the task will be the continuation from the previous year to witness the advances made by the community. More info.
Task 3.
Cells undergoing mitotic phase will express different amount of antibodies which would be a useful sign for narrowing down the possible patient pattern. Before determining the mitotic cells, the cells are first detected. As mentioned, the scientists need to ensure that there are at least one or two mitotic cells on each patient’s specimen. Due to their much lower number of occurrences, we name the second task as the mitotic cell detection. In this setting, we assume the interphase cells (i.e. the non-mitotic cells) as the “background” cells. Recently it was found that mitotic cell detection can addressed using the secondary counterstain. Again, the introduction of secondary counterstain may not be economical. Thus, in this work we only primarily aim on the problem of mitotic cell detection using the primary counterstain. More info.
Task 4.
Cell segmentation is the first step on the whole analysis. Despite its high difficulties, the segmentation problem can be solved by the secondary channel using DAPI. Nevertheless, the introduction of DAPI may complicate the laboratory work flow; thus, may not even practical in some pathology laboratories. This means, the cell segmentation on the primary channel is still not solved yet. The aim for this task is using only the primary channel, to generate the segmentation map that is equivalent to the map generated from the secondary channel. More info.